StackGP Parallel Demo#
This is a demo demonstrating the advantages of using parallel exectution with StackGP.
First we need to load in the necessary packages
import StackGP as sgp
import numpy as np
import random
Define Benchmarks#
Here we define 10 random benchmark problems using the StackGP generateRandomBenchmark function.
# define train/test split function
def trainTestSplit(inputData, response, testSize=0.2, randomState=None):
if randomState is not None:
np.random.seed(randomState)
indices = np.arange(len(inputData[0]))
np.random.shuffle(indices)
splitIndex = int(len(inputData[0]) * (1 - testSize))
trainIndices = indices[:splitIndex]
testIndices = indices[splitIndex:]
return inputData[:, trainIndices], response[trainIndices], inputData[:, testIndices], response[testIndices]
# Benchmark 1
# print seed for reproducibility
# generate random interger
np.random.seed(None) # reset seed to random
seed = np.random.randint(0, 1000)
print(f"Seed: {seed}")
np.random.seed(seed)
random.seed(seed)
inputData1, response1, model1 = sgp.generateRandomBenchmark(numVars=5, numSamples=100)
trainInput1, trainResponse1, testInput1, testResponse1 = trainTestSplit(inputData1, response1, testSize=0.2, randomState=1)
sgp.printGPModel(model1)
Seed: 683
# Benchmark 2
seed = 13
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData2, response2, model2 = sgp.generateRandomBenchmark(numVars=5, numSamples=100)
trainInput2, trainResponse2, testInput2, testResponse2 = trainTestSplit(inputData2, response2, testSize=0.2, randomState=2)
sgp.printGPModel(model2)
Seed: 13
# Benchmark 3
seed = 27
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData3, response3, model3 = sgp.generateRandomBenchmark(numVars=2, numSamples=100)
trainInput3, trainResponse3, testInput3, testResponse3 = trainTestSplit(inputData3, response3, testSize=0.2, randomState=3)
sgp.printGPModel(model3)
Seed: 27
# Benchmark 4
seed = 40
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData4, response4, model4 = sgp.generateRandomBenchmark(numVars=10, maxLength=20, numSamples=100)
trainInput4, trainResponse4, testInput4, testResponse4 = trainTestSplit(inputData4, response4, testSize=0.2, randomState=4)
sgp.printGPModel(model4)
Seed: 40
# Benchmark 5
seed = 100
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData5, response5, model5 = sgp.generateRandomBenchmark(numVars=12, maxLength=10, numSamples=40)
trainInput5, trainResponse5, testInput5, testResponse5 = trainTestSplit(inputData5, response5, testSize=0.2, randomState=5)
sgp.printGPModel(model5)
Seed: 100
# Benchmark 6
seed = 205
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData6, response6, model6 = sgp.generateRandomBenchmark(numVars=20, maxLength=15, numSamples=200)
trainInput6, trainResponse6, testInput6, testResponse6 = trainTestSplit(inputData6, response6, testSize=0.2, randomState=6)
sgp.printGPModel(model6)
Seed: 205
# Benchmark 7
seed = 1
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData7, response7, model7 = sgp.generateRandomBenchmark(numVars=15, maxLength=25, numSamples=150)
trainInput7, trainResponse7, testInput7, testResponse7 = trainTestSplit(inputData7, response7, testSize=0.2, randomState=7)
sgp.printGPModel(model7)
Seed: 1
# Benchmark 8
seed = 2
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData8, response8, model8 = sgp.generateRandomBenchmark(numVars=25, maxLength=30, numSamples=300)
trainInput8, trainResponse8, testInput8, testResponse8 = trainTestSplit(inputData8, response8, testSize=0.2, randomState=8)
sgp.printGPModel(model8)
Seed: 2
# Benchmark 9
seed = 3
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData9, response9, model9 = sgp.generateRandomBenchmark(numVars=30, maxLength=35, numSamples=400)
trainInput9, trainResponse9, testInput9, testResponse9 = trainTestSplit(inputData9, response9, testSize=0.2, randomState=9)
sgp.printGPModel(model9)
Seed: 3
# Benchmark 10
seed = 4
print(f"Seed: {seed}") # print seed for reproducibility
np.random.seed(seed)
random.seed(seed)
inputData10, response10, model10 = sgp.generateRandomBenchmark(numVars=5, maxLength=15, numSamples=500, opsChoices=sgp.allOps())
trainInput10, trainResponse10, testInput10, testResponse10 = trainTestSplit(inputData10, response10, testSize=0.2, randomState=10)
sgp.printGPModel(model10)
Seed: 4
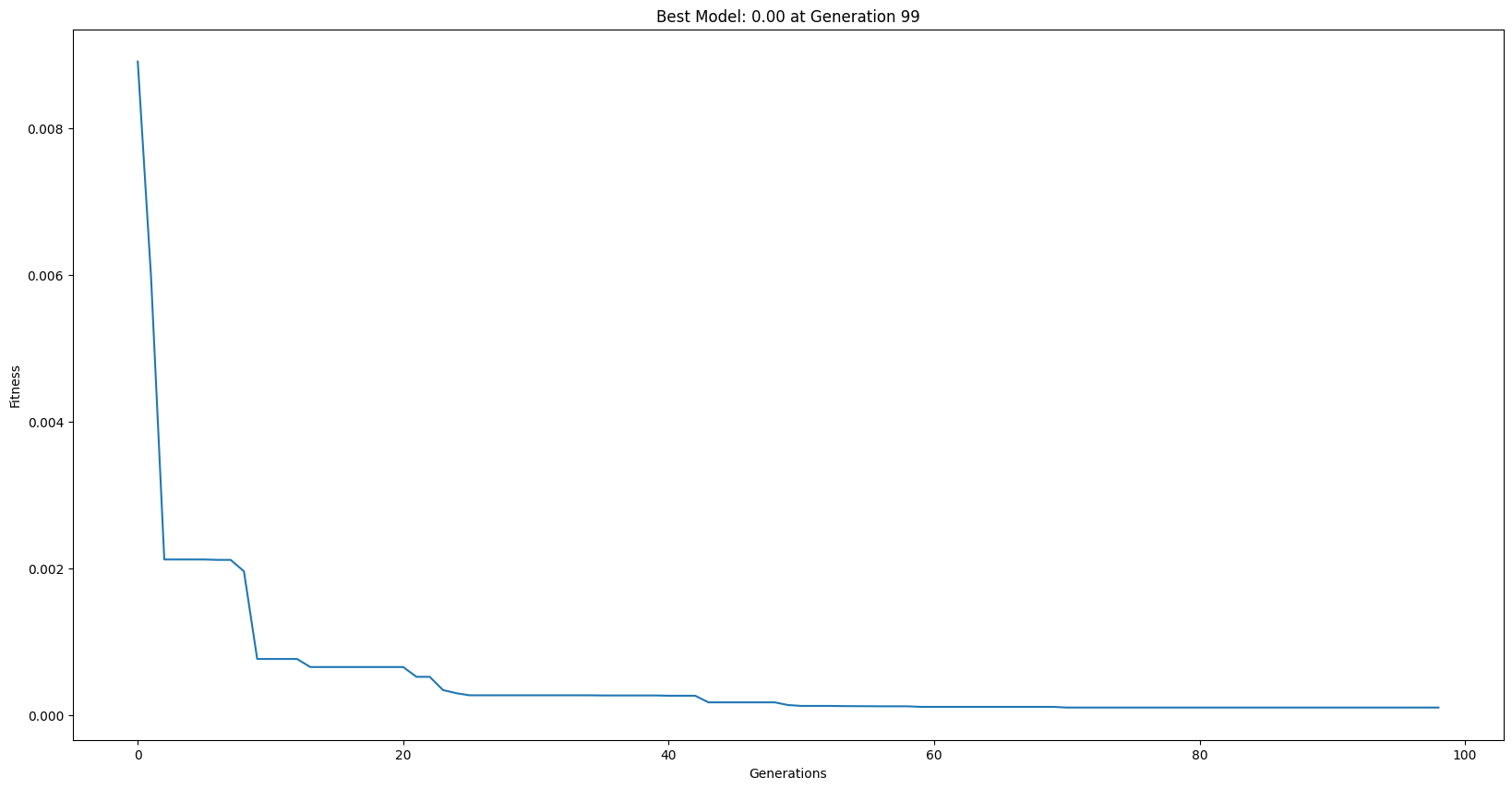
Example Serial Run#
_ = sgp.evolve(trainInput1, trainResponse1, liveTracking=True)
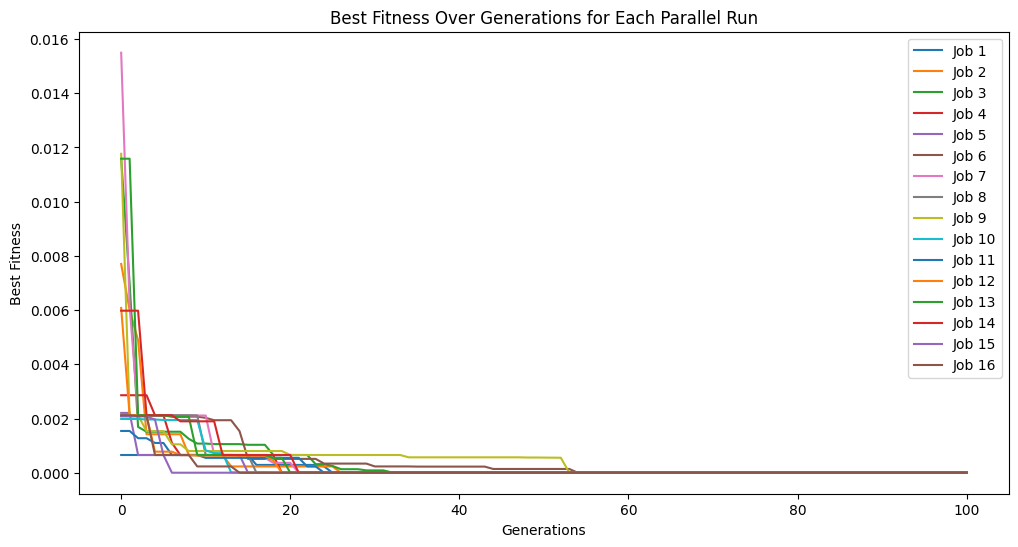
Example Parallel Run#
We can see the evolutionary traces from all of the parallel searches.
_ = sgp.parallelEvolve(trainInput1, trainResponse1, tracking=True)
Running parallel evolution with 16 jobs.
Run Evolutionary Searches (Serial)#
Now we run 10 serial searches for each problem (5 times for each benchmark). We will use 10 second runs for each search. From each run, we take the best model on the training set and evaluate using the test set.
# Benchmark 1 Train (Serial)
modelsSerial1 = [sgp.evolve(trainInput1, trainResponse1, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance1 = [sgp.fitness(model, testInput1, testResponse1) for model in modelsSerial1]
# Benchmark 2 Train (Serial)
modelsSerial2 = [sgp.evolve(trainInput2, trainResponse2, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance2 = [sgp.fitness(model, testInput2, testResponse2) for model in modelsSerial2]
# Benchmark 3 Train (Serial)
modelsSerial3 = [sgp.evolve(trainInput3, trainResponse3, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance3 = [sgp.fitness(model, testInput3, testResponse3) for model in modelsSerial3]
# Benchmark 4 Train (Serial)
modelsSerial4 = [sgp.evolve(trainInput4, trainResponse4, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance4 = [sgp.fitness(model, testInput4, testResponse4) for model in modelsSerial4]
# Benchmark 5 Train (Serial)
modelsSerial5 = [sgp.evolve(trainInput5, trainResponse5, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance5 = [sgp.fitness(model, testInput5, testResponse5) for model in modelsSerial5]
# Benchmark 6 Train (Serial)
modelsSerial6 = [sgp.evolve(trainInput6, trainResponse6, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance6 = [sgp.fitness(model, testInput6, testResponse6) for model in modelsSerial6]
# Benchmark 7 Train (Serial)
modelsSerial7 = [sgp.evolve(trainInput7, trainResponse7, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance7 = [sgp.fitness(model, testInput7, testResponse7) for model in modelsSerial7]
# Benchmark 8 Train (Serial)
modelsSerial8 = [sgp.evolve(trainInput8, trainResponse8, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance8 = [sgp.fitness(model, testInput8, testResponse8) for model in modelsSerial8]
# Benchmark 9 Train (Serial)
modelsSerial9 = [sgp.evolve(trainInput9, trainResponse9, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance9 = [sgp.fitness(model, testInput9, testResponse9) for model in modelsSerial9]
# Benchmark 10 Train (Serial)
modelsSerial10 = [sgp.evolve(trainInput10, trainResponse10, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
serialTestPerformance10 = [sgp.fitness(model, testInput10, testResponse10) for model in modelsSerial10]
Run Evolutionary Searches (Parallel)#
Here we run the same searches but this time use all parallel cores on the machine to parallelize the search. The same time constraints are utilized.
# Benchmark 1 Train (Parallel)
modelsParallel1 = [sgp.parallelEvolve(trainInput1, trainResponse1, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance1 = [sgp.fitness(model, testInput1, testResponse1) for model in modelsParallel1]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 2 Train (Parallel)
modelsParallel2 = [sgp.parallelEvolve(trainInput2, trainResponse2, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance2 = [sgp.fitness(model, testInput2, testResponse2) for model in modelsParallel2]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 3 Train (Parallel)
modelsParallel3 = [sgp.parallelEvolve(trainInput3, trainResponse3, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance3 = [sgp.fitness(model, testInput3, testResponse3) for model in modelsParallel3]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 4 Train (Parallel)
modelsParallel4 = [sgp.parallelEvolve(trainInput4, trainResponse4, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance4 = [sgp.fitness(model, testInput4, testResponse4) for model in modelsParallel4]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 5 Train (Parallel)
modelsParallel5 = [sgp.parallelEvolve(trainInput5, trainResponse5, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance5 = [sgp.fitness(model, testInput5, testResponse5) for model in modelsParallel5]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 6 Train (Parallel)
modelsParallel6 = [sgp.parallelEvolve(trainInput6, trainResponse6, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance6 = [sgp.fitness(model, testInput6, testResponse6) for model in modelsParallel6]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 7 Train (Parallel)
modelsParallel7 = [sgp.parallelEvolve(trainInput7, trainResponse7, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance7 = [sgp.fitness(model, testInput7, testResponse7) for model in modelsParallel7]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 8 Train (Parallel)
modelsParallel8 = [sgp.parallelEvolve(trainInput8, trainResponse8, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance8 = [sgp.fitness(model, testInput8, testResponse8) for model in modelsParallel8]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 9 Train (Parallel)
modelsParallel9 = [sgp.parallelEvolve(trainInput9, trainResponse9, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance9 = [sgp.fitness(model, testInput9, testResponse9) for model in modelsParallel9]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
# Benchmark 10 Train (Parallel)
modelsParallel10 = [sgp.parallelEvolve(trainInput10, trainResponse10, generations=1000, capTime=True, timeLimit=10)[0] for _ in range(5)]
parallelTestPerformance10 = [sgp.fitness(model, testInput10, testResponse10) for model in modelsParallel10]
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
Running parallel evolution with 16 jobs.
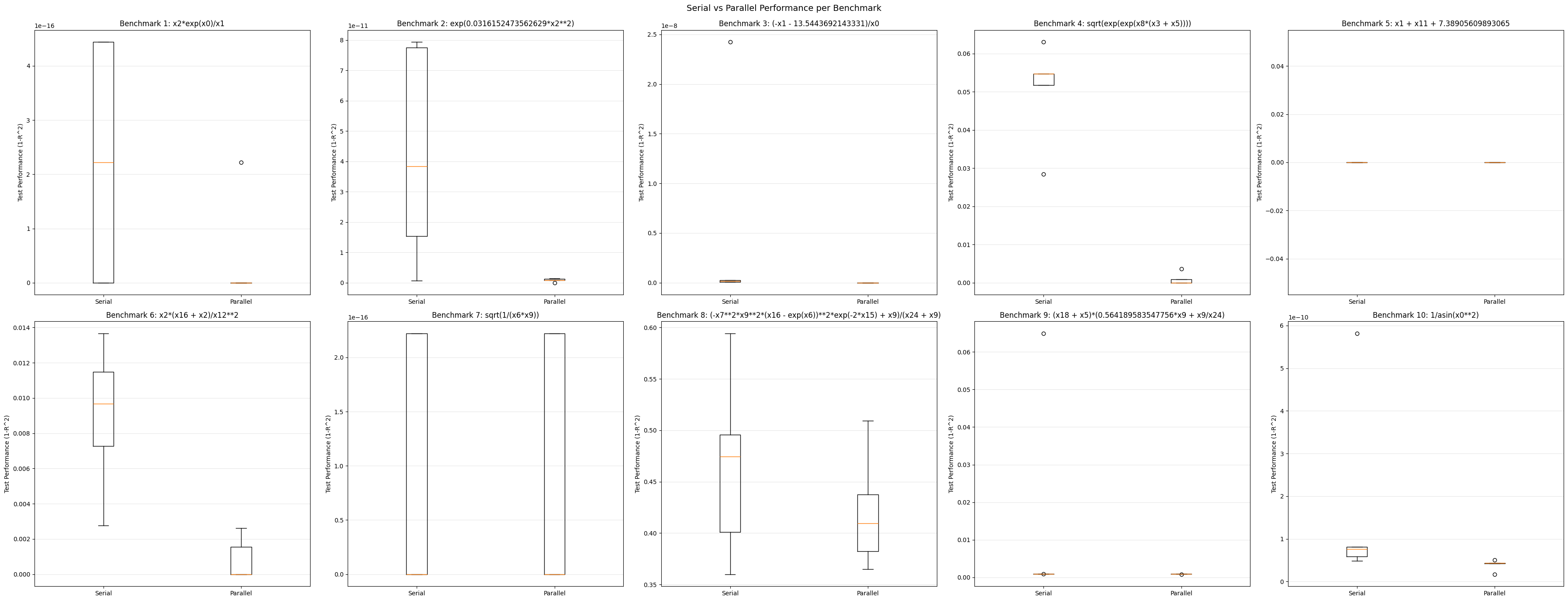
Results#
Now we can compare the performance of the two approaches across all 10 benchmark problems.
fig, axes = plt.subplots(2, 5, figsize=(18*2, 2*7), sharey=False)
benchmarkModels = [model1, model2, model3, model4, model5, model6, model7, model8, model9, model10]
for i, ax in enumerate(axes.ravel()):
ax.boxplot(
[serialPerformances[i], parallelPerformances[i]],
labels=["Serial", "Parallel"]
)
# set y label
ax.set_ylabel("Test Performance (1-R^2)")
ax.set_title(f"Benchmark {i+1}: {sgp.printGPModel(benchmarkModels[i])}")
ax.grid(axis="y", alpha=0.3)
fig.suptitle("Serial vs Parallel Performance per Benchmark", fontsize=14)
plt.tight_layout()
plt.savefig('/Users/nathanhaut/Downloads/serial_parallel_comparison.jpeg', format='jpeg')
plt.show()
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
/var/folders/tn/s_jbjrf525zd165vpvwykqk40000gn/T/ipykernel_72835/2947987728.py:4: MatplotlibDeprecationWarning: The 'labels' parameter of boxplot() has been renamed 'tick_labels' since Matplotlib 3.9; support for the old name will be dropped in 3.11.
ax.boxplot(
serialPerformances = [serialTestPerformance1, serialTestPerformance2, serialTestPerformance3, serialTestPerformance4, serialTestPerformance5, serialTestPerformance6, serialTestPerformance7, serialTestPerformance8, serialTestPerformance9, serialTestPerformance10]
parallelPerformances = [parallelTestPerformance1, parallelTestPerformance2, parallelTestPerformance3, parallelTestPerformance4, parallelTestPerformance5, parallelTestPerformance6, parallelTestPerformance7, parallelTestPerformance8, parallelTestPerformance9, parallelTestPerformance10]
# Compute average performance improvement from each serial case to each parallel case across the 10 benchmarks
averageImprovements = []
for i in range(10):
averageImprovements.append([])
for j in range(5):
improvement = (serialPerformances[i][j]-parallelPerformances[i][j])/abs(serialPerformances[i][j])
averageImprovements[i].append(improvement)
print("Average Performance Improvements from Serial to Parallel Across Benchmarks:")
for i, benchmarkImprovements in enumerate(averageImprovements):
perfMean = np.median(benchmarkImprovements)
# if not nan
if not np.isnan(perfMean):
print(f"Benchmark {i+1}: {perfMean:.4f}")
else:
print(f"Benchmark {i+1}: NaN (No improvements, solved)")
Average Performance Improvements from Serial to Parallel Across Benchmarks:
Benchmark 1: NaN (No improvements, solved)
Benchmark 2: 0.9633
Benchmark 3: 0.9998
Benchmark 4: 1.0000
Benchmark 5: NaN (No improvements, solved)
Benchmark 6: 1.0000
Benchmark 7: NaN (No improvements, solved)
Benchmark 8: 0.0899
Benchmark 9: 0.0000
Benchmark 10: 0.4459
Here we look at the median performance improvement across all runs.
# remove nans from averageImprovements
averageImprovementsClean = []
for i in range(10):
averageImprovementsClean.append([x for x in averageImprovements[i] if not np.isnan(x)])
# median of all values in averageImprovementsClean
np.median([item for sublist in averageImprovementsClean for item in sublist])
np.float64(0.9390946224295529)
Scaling Study#
Here we explore the size of the search space explored across varying numbers of cores. The code below modifies the parallelEvolve functionn so we can get the number of generations for the scaling study plots.
from joblib import Parallel, delayed
from StackGP import evolve, sortModels
import os
def parallelEvolveScaling(*args,n_jobs=-1,avail_cores=-1, **kwargs):
if avail_cores==-1:
try:
avail_cores=len(os.sched_getaffinity(0))
except:
avail_cores=os.cpu_count()
if n_jobs==-1:
try:
n_jobs=len(os.sched_getaffinity(0))
except:
n_jobs=os.cpu_count()
if "tracking" in kwargs and kwargs["tracking"]:
kwargs["returnTracking"]=True
print(f"Running parallel evolution with {n_jobs} jobs.")
if "liveTracking" in kwargs and kwargs["liveTracking"]:
print("Live tracking is not supported in parallel evolution, disabling live tracking.")
kwargs["liveTracking"]=False
runs = Parallel(n_jobs=avail_cores, backend="loky")(delayed(evolve)(*args, **kwargs) for _ in range(n_jobs))
if ("tracking" in kwargs and kwargs["tracking"]):
runs, tracking = zip(*runs)
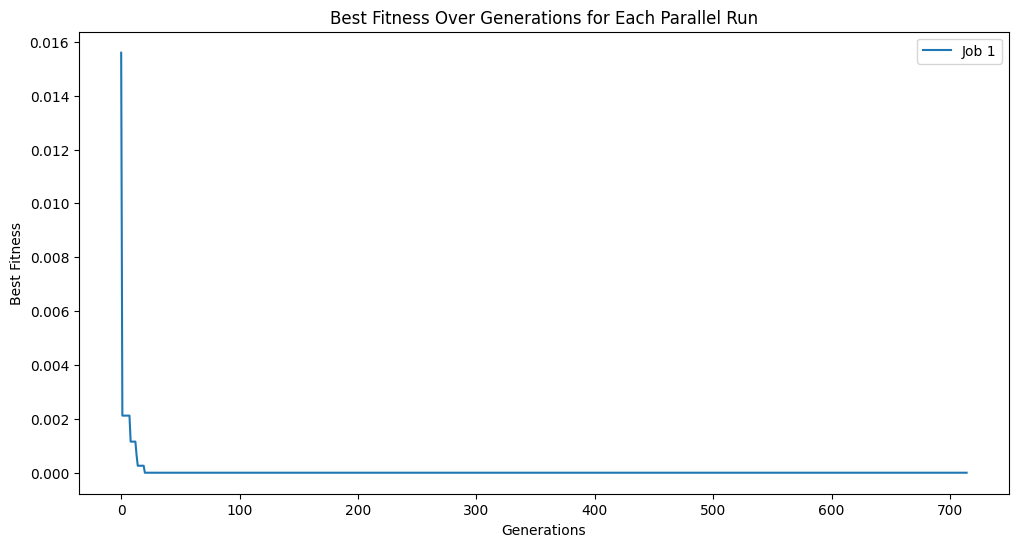
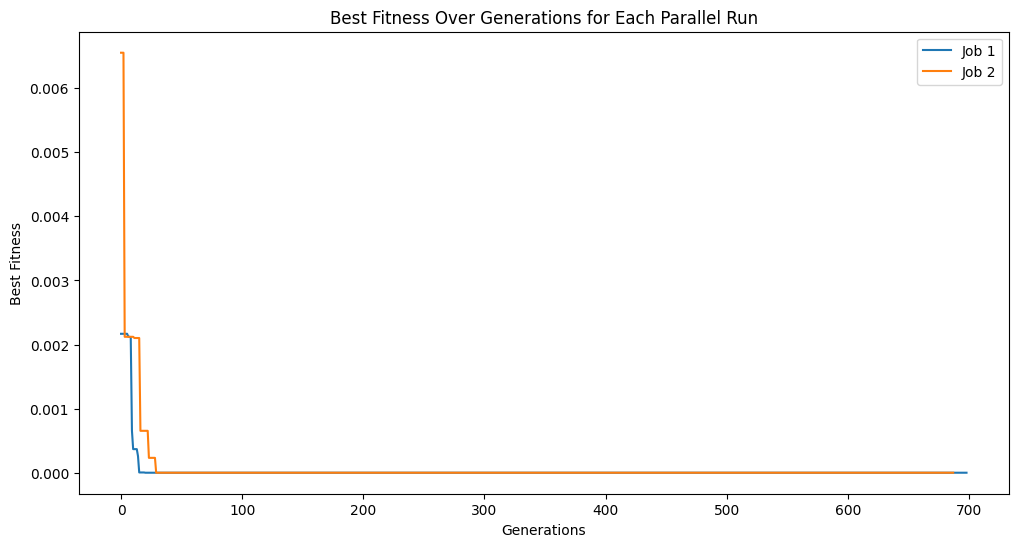
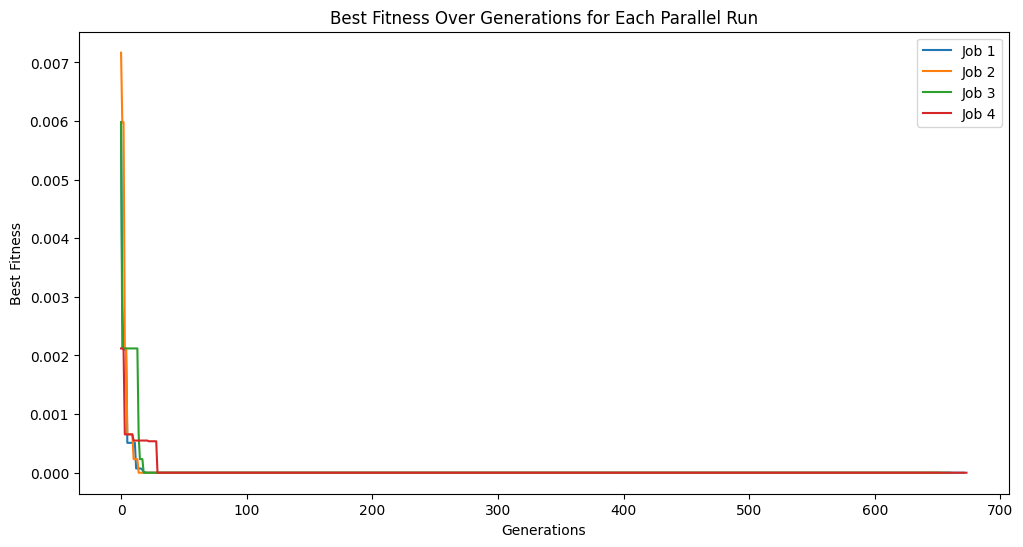
# plot tracking for each job
plt.figure(figsize=(12, 6))
for i, track in enumerate(tracking):
plt.plot(track, label=f'Job {i+1}')
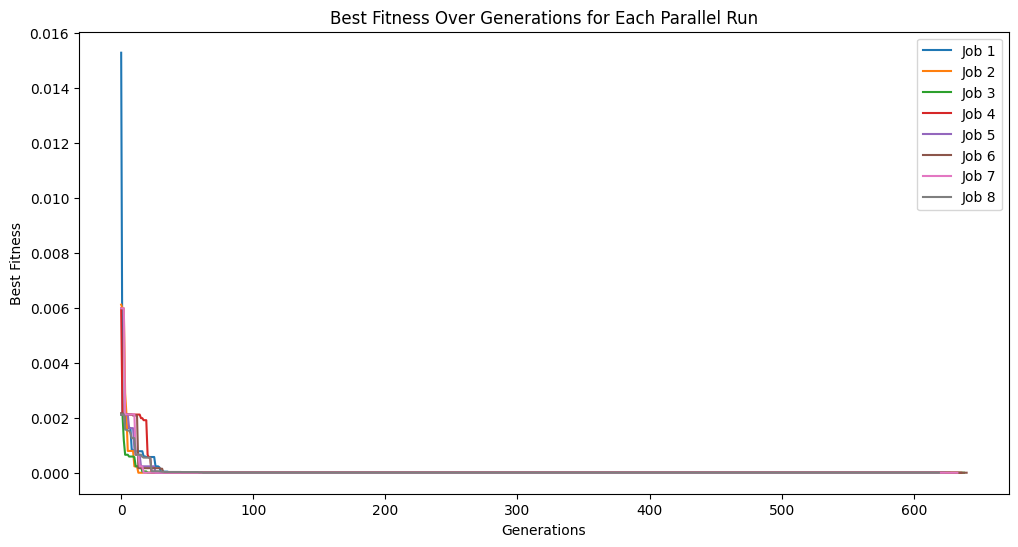
plt.title('Best Fitness Over Generations for Each Parallel Run')
plt.xlabel('Generations')
plt.ylabel('Best Fitness')
if n_jobs <= 16: # Only show legend if there are a reasonable number of jobs
plt.legend()
plt.show()
flat = [model for sublist in runs for model in sublist]
return sortModels(flat), tracking if ("tracking" in kwargs and kwargs["tracking"]) else None
trackingOutputs = []
for i in [1, 2, 4, 8, 16]:
trackingOutputs.append(parallelEvolveScaling(trainInput1, trainResponse1, generations=1000, capTime=True, timeLimit=10, n_jobs=i, tracking=True)[1])
Running parallel evolution with 1 jobs.
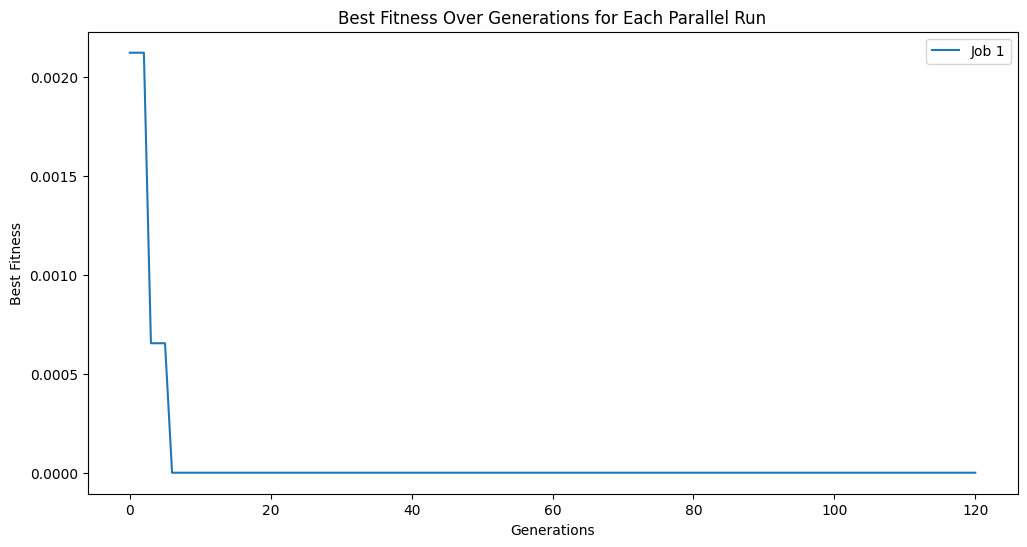
Running parallel evolution with 2 jobs.
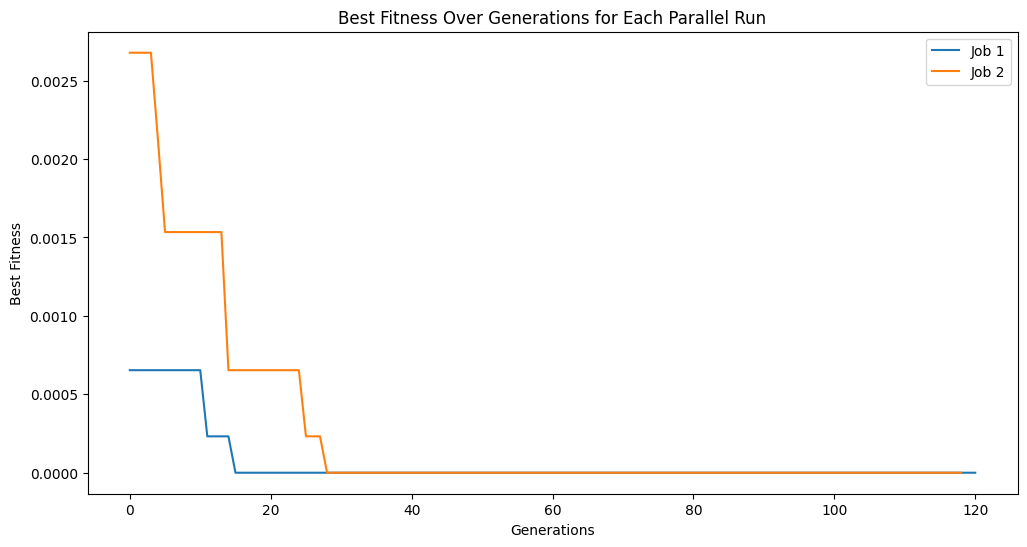
Running parallel evolution with 4 jobs.
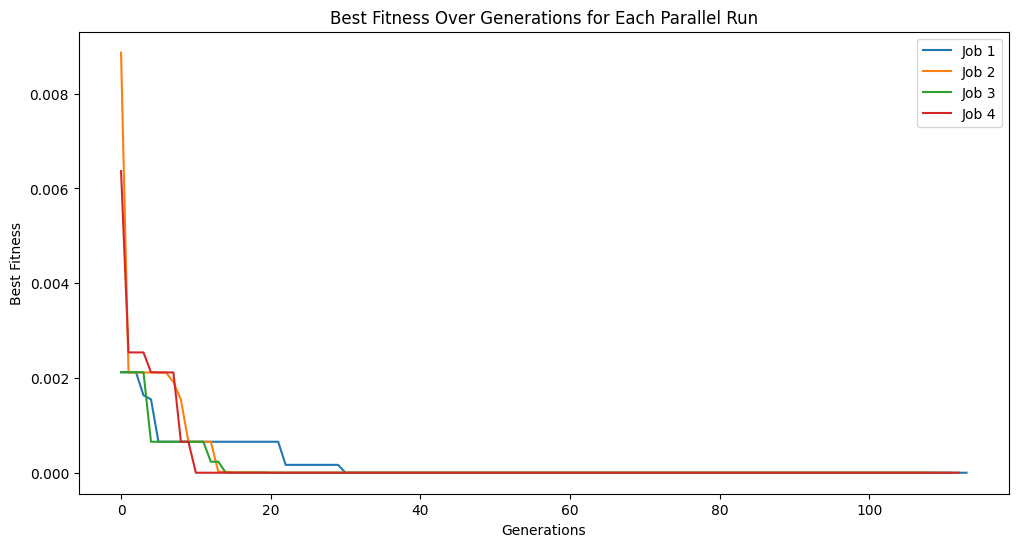
Running parallel evolution with 8 jobs.
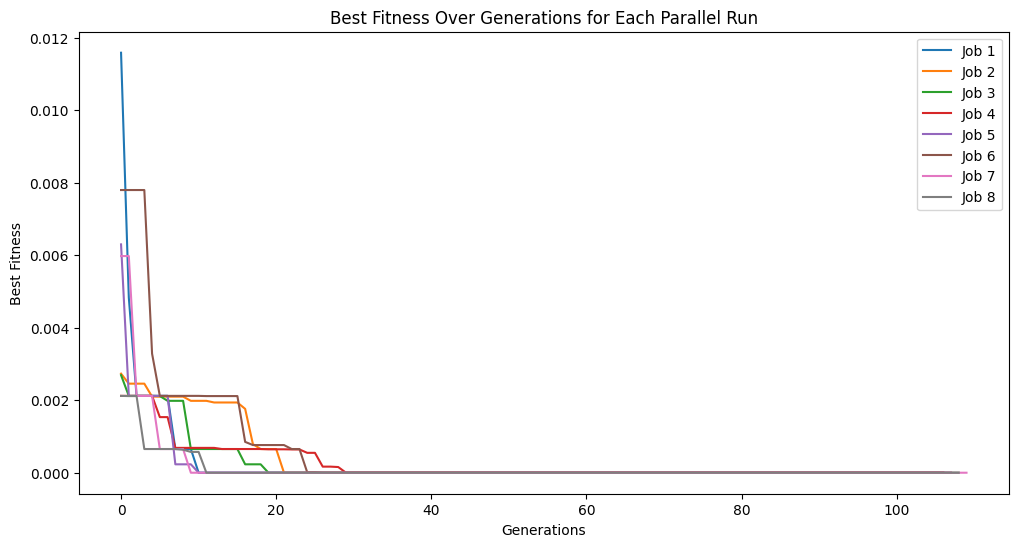
Running parallel evolution with 16 jobs.
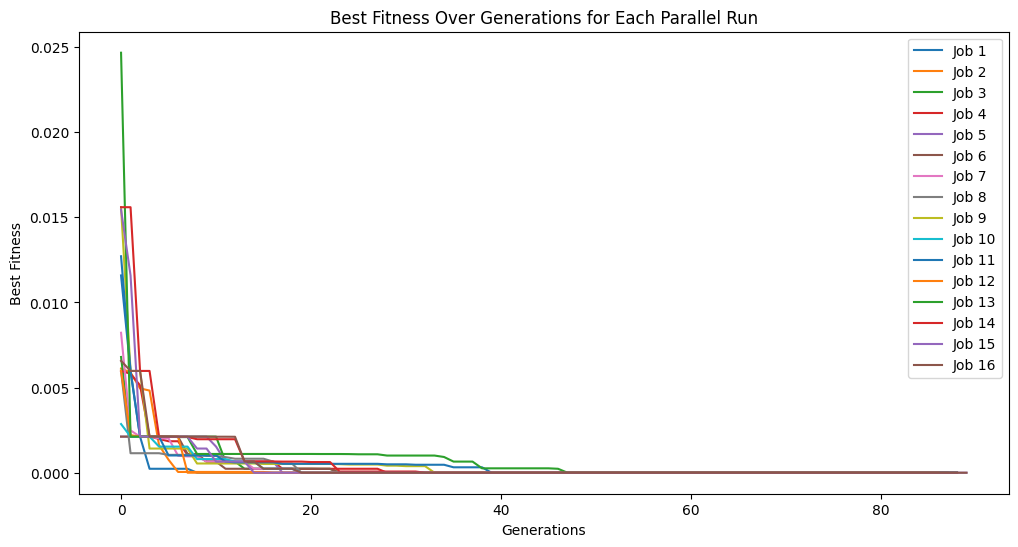
totalEvals = []
for track in trackingOutputs:
totalEvals.append(np.sum([len(gen) for gen in track]))
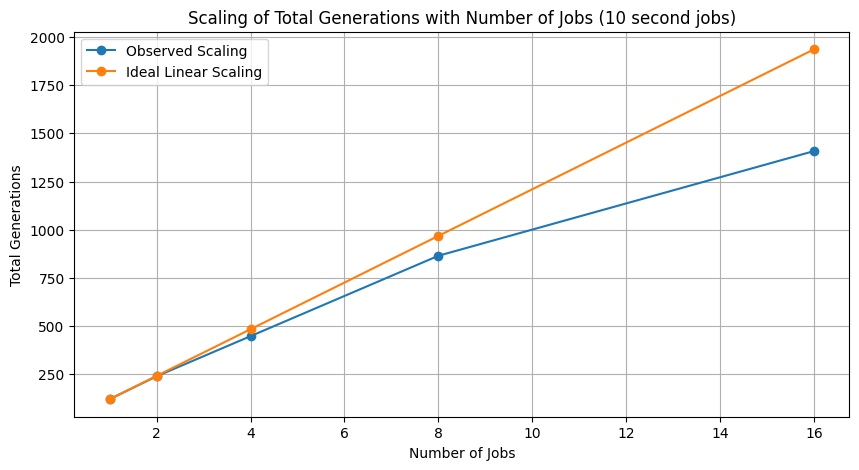
# plot scaling of total evaluations with number of jobs
plt.figure(figsize=(10, 5))
plt.plot([1, 2, 4, 8, 16], totalEvals, marker='o', label="Observed Scaling")
plt.plot([1, 2, 4, 8, 16], [totalEvals[0] * i for i in [1, 2, 4, 8, 16]], marker='o', label='Ideal Linear Scaling')
#plt.xscale('log', base=2)
plt.xlabel('Number of Jobs')
plt.ylabel('Total Generations')
plt.title('Scaling of Total Generations with Number of Jobs (10 second jobs)')
plt.legend()
plt.grid()
# export plot as jpeg
plt.savefig('/Users/nathanhaut/Downloads/scaling_plot.jpeg', format='jpeg')
plt.show()
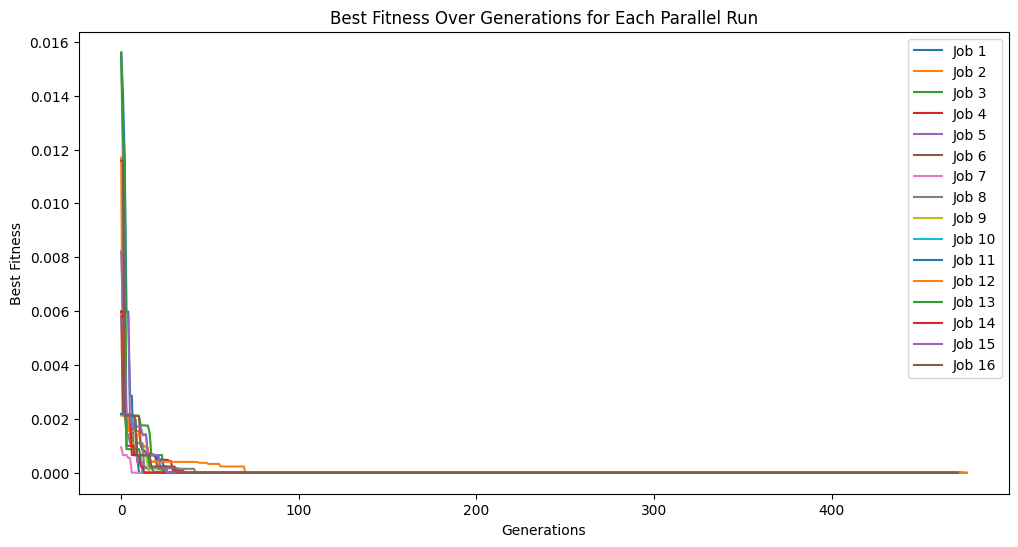
trackingOutputs1Min = []
for i in [1, 2, 4, 8, 16]:
trackingOutputs1Min.append(parallelEvolveScaling(trainInput1, trainResponse1, generations=1000, capTime=True, timeLimit=60, n_jobs=i, tracking=True)[1])
Running parallel evolution with 1 jobs.
Running parallel evolution with 2 jobs.
Running parallel evolution with 4 jobs.
Running parallel evolution with 8 jobs.
Running parallel evolution with 16 jobs.
totalEvals1Min = []
for track in trackingOutputs1Min:
totalEvals1Min.append(np.sum([len(gen) for gen in track]))
# plot scaling of total evaluations with number of jobs
plt.figure(figsize=(10, 5))
plt.plot([1, 2, 4, 8, 16], totalEvals1Min, marker='o', label="Observed Scaling")
plt.plot([1, 2, 4, 8, 16], [totalEvals1Min[0] * i for i in [1, 2, 4, 8, 16]], marker='o', label='Ideal Linear Scaling')
#plt.xscale('log', base=2)
plt.xlabel('Number of Jobs')
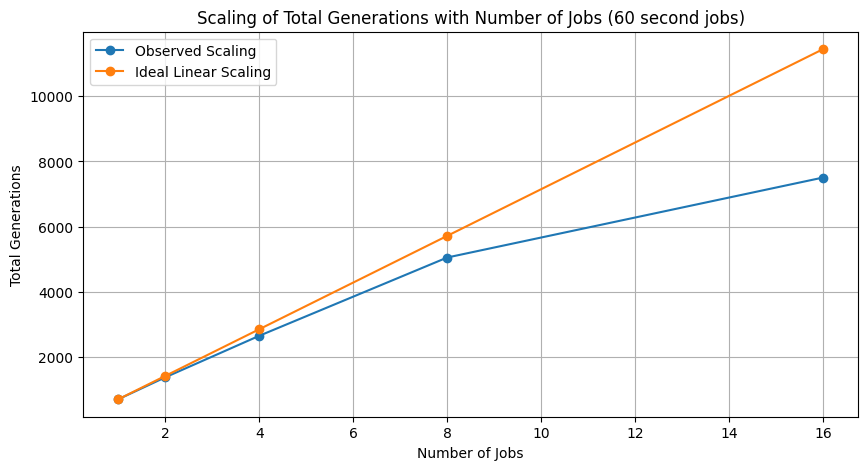
plt.ylabel('Total Generations')
plt.title('Scaling of Total Generations with Number of Jobs (60 second jobs)')
plt.legend()
plt.grid()
# export plot as jpeg
plt.savefig('/Users/nathanhaut/Downloads/scaling_plot_1min.jpeg', format='jpeg')
plt.show()
Performance improvement using 16 cores for 10 second case.
totalEvals[-1]/totalEvals[0]
np.float64(11.636363636363637)
Performance improvement using 16 cores for 1 minute case.
totalEvals1Min[-1]/totalEvals1Min[0]
np.float64(10.499300699300699)